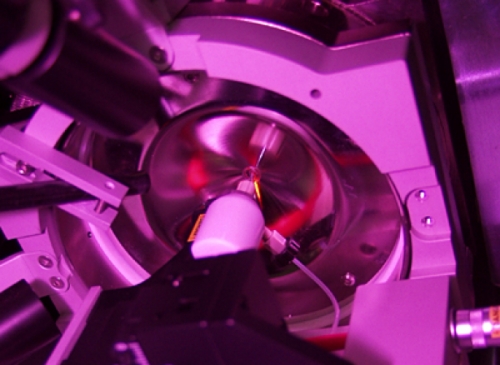
You’ve heard all about DNA. It’s sort of like your body’s musical score.
But what actually makes the music? Proteins. Inside your cells, as many as a million proteins may be at work — making things happen, folding into different structures, changing each other.
These proteins work together to create a normally-functioning cell much like the sounds from different instruments come together to create a well-played piece of music.
For scientists, figuring out an organism’s proteome is like taking apart a symphony. Which instrument is responsible for that high note? What combination of sounds produces perfect harmony?
Your genes, fixed at birth, could be called your body’s musical score. The DNA code instructs the body to make various proteins. But deep inside your cells, those proteins are carrying out your body’s functions — making its music. “Proteins are the molecular machines that actually do the job,” says Alex Tropsha, associate professor of pharmacy. Proteins fold into various structures, act upon each other, change in response to other parts of the cell. Proteins are so numerous and so busy that figuring out an organism’s proteome — the properties and activities of each of its proteins — promises to be thousands of times more complicated than figuring out its genome.
Proteomics, as this new field is called, emerged only in the last five years, after scientists sequenced the genomes of humans, fruit flies, yeast, and other creatures. Proteomics wouldn’t be possible without those genome definitions. An essential technique of proteomics — mass spectrometry — basically helps identify proteins by smashing them up and looking at their pieces. It separates proteins into charged particles, then “weighs” those particles. The weight, or mass, of each particle yields a “fingerprint” that a computer can match to a database of amino acids, which are the building blocks of proteins. The amino acid information can be matched to all known gene sequences to identify the protein. “You couldn’t do proteomics without the genome sequences,” says Bill Marzluff, professor of biochemistry. “That’s why proteomics has become important recently.”
The more familiar field of genomics and its burgeoning offshoot, proteomics, are helping scientists learn about how cells carry on the business of our bodies. But the more these two fields tell us, the more questions we have. Probably less than 50 percent of the time do genomics and proteomics agree, says Lee Graves, associate professor of pharmacology. For instance, some studies show that while levels of mRNA (an intermediate stage between DNA and protein) increase, the amount of protein produced actually decreases. So just because a gene codes for a protein doesn’t necessarily mean that the protein gets made. Scientists hope that as the field of proteomics grows, it will provide more answers. “Proteomics allows us to skip over the complexity of gene regulation and look directly at changes in the proteins,” Graves says. “This is one of the reasons why proteomics is now so popular.”
Carolina is getting into the proteomics field while it’s new, thanks to an anonymous $25 million donation in honor of the late Chancellor Michael Hooker. The gift funded a new proteomics facility and equipment. “It has allowed all of us here to do experiments and accomplish things we couldn’t have done before,” Marzluff says. “Proteomics means a new tool to try to solve problems that people have been working on for twenty years in some cases.”
For example, Richard Boucher, director of Carolina’s Cystic Fibrosis Center, is applying proteomics to understand the protein that is known to be defective in Cystic Fibrosis (see Mapping Disease). And Jackson Stutts, associate professor of medicine, is leading a team that received a $1.79 million grant from the Cystic Fibrosis Foundation to apply proteomics to understanding the disease. Carolina departments involved in proteomics research include nearly all the departments in the medical and health sciences as well as chemistry and biology.
Proteomics promises to yield great insight into the workings of our bodies, but that insight won’t come easy. Because proteins don’t work alone, analyzing them will require digesting an amazing overload of information. Small modifications that happen to proteins can mean big changes in function. The addition of chemical groups such as phosphates or methyl groups, for instance, can be required for a protein to function, or they can make another protein stop working. Proteins can also modify each other. The possible combinations and outcomes boggle the mind.
Analyzing, storing, and retrieving these vast amounts of information will take some high-tech tools. At Carolina, Christoph Borchers, faculty director of Carolina’s Proteomics Core Facility, is the keeper of those tools. Scientists perfecting proteomics technology are working toward achieving “high throughput” — analyzing as many proteins as fast as possible. Each week Borchers is getting new modifications for the facility; by the time you’re reading this article, the equipment will be able to analyze 9,600 samples at once. Carolina is one of five U.S.“validation sites” that are testing some of the newest equipment before it’s made available commercially.
But proteomics technology in general still has some growing to do. Compare the rate of proteomics analysis — 9,600 samples at once — to the fastest genomics analysis — 20,000 samples. Proteins present so many more complications than genes that the technology has yet to catch up.
Some researchers believe that combining genomics and proteomics will yield the best results. Scientists at UNC’s Center for Genomic Sciences are beginning to collaborate with scientists using proteomics tools such as crystallography and protein modeling to help guide their work, says Terry Magnuson, director of the center. The classic way of learning about genes’ functions is to make a sequence variation — commonly called a mutation — in a particular gene and then observe the effect on say, a mouse or a fly. Magnuson says, “We’d like to get to the point where if we make a mutation in a gene, we can ask the question ahead of time, ‘what do we think that mutation is going to do to that protein structure, and do we even want to make a mouse out of it?’”
Tropsha and John Sondek, assistant professor of pharmacology, help Magnuson answer that question using two different approaches to studying protein structure. The way that proteins fold in on themselves, the shapes and patterns they make, often determines protein function, and learning more about how folding happens can help scientists find proteins that would make good targets for drugs. Tropsha uses computer modeling to predict protein structure. Sondek takes the experimental route, using crystallography to actually examine and see a protein’s structure.
Charles Perou, assistant professor of genetics, is also beginning to combine genetics with proteomics. In studying breast cancer tumors, Perou uses microscopy and mRNA to create images known as microarrays, which show 20,000 fluorescent spots representing the expression of as many as 19,000 genes. Perou uses the microarrays to help him classify tumors into various groups. “We had twenty patients with tumors that were all lumped into one group, and the microarray information has helped us divide those tumors into five different groups,” he says. Perou knows, for example, that one group of tumors is resistant to treatment and presents a poor prognosis, while another group responds well to treatment.
Now Perou is beginning to work with Borchers to get similar information about proteins involved in these tumors to help define the groups even further, with the hope of developing specific treatments for each group. Marzluff says, “There aren’t too many places that can say they can do both microarrays and proteomics real well right now.”
Another area that may benefit from proteomics: mouse genetics. Marzluff says, “With all the mouse genetics we have here, as the mouse proteome becomes available for analysis we would be in a real position to become even more of a leader in mouse biology.” And if Carolina stays in the lead of the high-throughput race, he adds, “I think we could easily become one of the leading proteomics centers in the country.”
An Orchestra of Proteins
Imagine listening to multiple pieces of music simultaneously and trying to identify each instrument and the part that it’s playing. Proteomics researchers face a similar challenge as they struggle to determine the role of each protein in the body.
Proteomics researchers work to determine what proteins are present in a cell, where they are located, how much of each protein is present, and how the proteins function. But because cells are dynamic, the protein constitution of a single cell is constantly changing, so some proteins may not even be present at certain stages of the cell cycle. If you add to this the fact that proteins are continuously modified, the task proteomics seeks to accomplish seems insurmountable.
But Christoph Borchers, assistant professor of biochemistry and biophysics and faculty director of Carolina’s Proteomics Core Facility, believes that mass spectrometry (MS) is just the technology for the job. He and his colleagues have assembled a state-of-the-art automated facility, offering the latest way to look at the entire protein environment of a cell. “With this technique you can listen to an entire orchestra of proteins,” Borchers says.
To understand Borchers’ excitement, you need to understand MS. First, a molecule is vaporized and charged, or ionized. Until 5 to 10 years ago, ionization and vaporization of fragile molecules were the Achilles heels of biological MS. Traditional techniques used heat, which could destroy peptides or proteins. But today, gentle ionization and vaporization techniques can safely transport charged peptides and proteins into the gas phase.
The mass spectrometer then measures each ion’s time of flight — the time it takes for the ion to reach the detector. The time of flight is affected by both an ion’s charge and its weight or mass. For instance, ions with lower weights will travel faster and will have a shorter time of flight. So the time of flight is used to determine each ion’s mass-to-charge ratio, which scientists can then use to identify the peptide or protein in question.
Borchers explains that MS has three distinct advantages over existing technology. MS is extremely accurate, able to distinguish a protein that weighs 1000.03 Daltons (a Dalton is the unit for protein weight) from one that weighs 1000.04 Daltons. MS is also extremely sensitive, requiring very little protein material. One of the three instruments at the Proteomics Core Facility is able to detect a femtamole of protein — a feat similar to detecting the addition of one drop of water to your backyard pool. This level of sensitivity allows researchers to use the native protein from cells instead of synthetic protein.
Finally, MS is able to provide sequence data that can be used to identify unknown proteins. Identifying all of the amino acids and their order in a given protein is referred to as protein sequencing. This process starts with digestion of the protein with an enzyme that systematically chops up the protein into smaller peptide fragments. The molecular weights of the peptides are then measured by MS so accurately that the protein can be identified by searching these masses against a protein or genome database. Like the unique grooves and coloring of jigsaw pieces, the masses of amino acid fragments indicate how the fragments can be pieced together to reveal the whole protein.
Borchers is also using MS to characterize proteins. “I call it proteomics, the second generation,” he says, noting that “protein characterization is still not easy.” Protein modifications such as phosphorylation and glycosylation attach additional chemical groups onto the protein, increasing its weight — but not by much. Low-weight modifications are difficult to detect by mass spectrometry. And, many of the modifications result in a negative charge on the protein, also making it difficult to detect.
“What proteomics needs to do, finally, is identify not only expressed proteins, but also identify and characterize altered proteins,” Borchers says. A collaboration with the Lineberger Comprehensive Cancer Center seeks to work on some of those questions. “What we want to do is characterize the proteins of the human breast cancer cell,” Borchers says. At 25,000 to 30,000 proteins, the breast cancer cell will certainly be a proving ground for proteomics.
Uncharted Waters
If it weren’t for Carolina’s commitment to proteomics, Michael Giddings, assistant professor of microbiology and immunology and of biomedical engineering, might not have come to Carolina.
Giddings’ postdoctoral work at the University of Utah got him thinking about the problems of current proteomics methods. “Most people assume that if you can identify the gene that encodes the protein, then the work is done. But there are many things that can happen that alter protein production. The relationship between gene and protein is not linear,” Giddings says.
The Giddings lab develops computer software that will gather, calculate, and analyze data. “These data will potentially allow us to trace the entire pathway from protein to gene,” Giddings says. “This would give a better biochemical picture of how proteins are derived from the genome and also provide more information about the functional role of these proteins.”
One program — “Proclaim,” developed by lab technician Mark Holmes — uses the molecular weight of a protein, generated by mass spectrometry, to search a set of possible modifications that will allow this protein to match up with a protein mass listed in a database of such masses. The Giddings lab is also building a system that can analyze multiple kinds of mass spectrometry measurements and plug these into a data-analysis system. This Protein Inference Engine (PIE) will use the data to map back to the gene to figure out what pathways are used and when. In collaboration with Janne Cannon, professor of microbiology, Giddings is beginning to use this new technology to map the human pathogen Neisseria gonorrhoeae. This organism causes the sexually transmitted disease gonorrhea and billions of dollars in health care costs. N. gonorrhoeae produces variable surface proteins that interact with a host’s cells. According to Giddings, this “barrage of different looks” helps N. gonorrhoeae evade the body’s natural defenses.
“Understanding how this organism works will lead to new drugs and treatments that will reduce human suffering,” Giddings says. The system could also be applied to other bacteria and agents that could be used in bioterrorism. In addition, Giddings collaborates with experts studying cystic fibrosis and developmental psychologists in the study of juvenile behavior.
Giddings uses high-performance liquid chromatography (HPLC) to simplify the protein mixes that feed into mass spectrometry. Giddings’ group is one of only a few labs that look at intact proteins first. They also use enzymes that cut the protein into smaller pieces. The process produces a banding pattern called a peptide mass fingerprint that is specific to each protein. Giddings hopes to eventually take one of the peptide mass fingerprints and scan the human genome database to figure out what part of the genome might have expressed this pattern.
Giddings has high hopes for proteomics and sees it as one of a set of integrated technologies that analyzes a cell in its entirety. Now scientists can model only small portions of a cell, but eventually they will be able to model a complete cell using supercomputers. Giddings says, “As quickly as computer technology is advancing, it’s likely that we’ll be able to model at least some of the simpler cells in maybe five to ten years.”
Where the Protein Leads
Science has a way of leading researchers down different paths. Carol Otey, assistant professor of cell and molecular physiology, studies cell adhesion and motility and the pathways that regulate the cell shape. Her work is an excellent example of “following where the science leads.”
As a postdoctoral fellow in Keith Burridge’s lab, Otey discovered a protein that plays a key role in organizing the actin cytoskeleton. Actin is an abundant cellular protein that forms the filaments that give the cell its shape. This new protein, which she named palladin, is involved in actin assembly. “There’s no evidence that palladin binds to actin directly. Instead, it binds to multiple things that bind to actin,” Otey says.
Otey studies palladin function in fibroblasts (cells that give rise to connective tissue), neurons (nerve cells), and glia (support cells for neurons). When there is a cellular signal to change shape, palladin protein levels increase. There are a number of situations in which a cell will change shape. For example, tumor cells grow uncontrollably, but within the tumor there is a subpopulation of traveling tumor cells called metastatic cells, which have a different shape. This shape change involves the actin cytoskeleton and palladin. “If we can find a way to specifically interfere with metastasis, cancer would be a treatable disease,” Otey says. “We would like to see whether there is a difference in palladin in normal, tumor, and metastatic cells.”
In addition to understanding cancer, Otey wants to know what goes wrong when there is an injury to the brain or spinal cord. After a severe injury to the central nervous system (CNS) — the brain and spinal cord — the body experiences a permanent loss of nerve function, but after an injury to the peripheral nervous system (such as a cut in the skin, severing a sensory nerve), the nerves will recover.
Why do nerves in the peripheral nervous system recover better than those in the CNS? One explanation is that when there is an injury to the CNS, neurons are cut and star-shaped glia cells called astrocytes migrate to the area and form a net around the site of injury. If the neuron survives the injury, it is unable to create new connections because it can’t punch through the astrocytes. This obstacle is called a glial scar and is a phenomenon of the CNS rather than the peripheral nervous system. This is why the CNS doesn’t heal well. “The glial scar involves astrocyte motility and shape change, but the molecular events that control glial scar formation are not understood. If we could understand how these changes arise, we could possibly manipulate the scar and prevent them from forming,” Otey says.
Otey’s lab uses standard cell biology techniques to manipulate palladin expression in cell culture. She can artificially add palladin by introducing its DNA into cells or inhibit palladin by introducing antisense palladin DNA, which blocks the production of palladin. Using fluorescence microscopy or video microscopy, Otey can observe how the actin cytoskeleton is organized. Otey’s observations show that in normal cells, three hours after an injury, palladin levels increase and cluster along the injury site. Also, cells lacking palladin lose their shape.
Because palladin appears to play an important role in metastatic cell movement and glial scar formation, these studies have far-reaching implications for cancer and spinal cord injury research.
Histone Tales
Repetition breeds familiarity. In only a few short years, images of the spiraling and twisted DNA double helix have become firmly established as the iconic darling for the genome sciences, much like the candy-striped red and white pole had been to men’s barbershops everywhere.
Yes, one might say the double helix does contain an almost kitschy purity, evocative of a landmark scientific achievement few of us fully comprehend. Still, the image of the double helix remains only a representation of the molecule. “It would surprise a lot of people to know that DNA is not just two linear strands. It’s wrapped around histone proteins to form a highly folded complex called chromatin,” says Brian Strahl, assistant professor of biochemistry and biophysics. This complex of nucleic acids and proteins binds DNA into higher-order structures, ultimately forming a chromosome.
The core histones appear in all organisms that have nucleated cells, including yeast and mammals. Four core histone proteins (H2A, H2B, H3, H4) each contain a “head,” or globular domain, and an amino “tail.” Of interest to Strahl is that these histones, specifically processes that modify them, are thought to play a major role in controlling gene expression and cell division.
Another image. Think of chromatin’s structure as a telephone cord with a bead between each coil. Each bead represents a nucleosome — chromatin’s fundamental repeating unit consisting of DNA wrapped twice around the four histone “core” proteins, their tails wagging and sometimes touching outside the nucleosome. Meanwhile, a fifth histone (H1) serves as a “linker histone” between nucleosomes. Now fold the cord on itself again and again.
This image approximates the scientifically known. Stretches of nucleosomes are folded upon themselves to create higher-order chromatin structures, albeit still not well defined. Although the chromatin packaging allows efficient storage of genetic information (the length of the entire complement of 46 chromosomes in a human cell is about one meter), it also impedes a wide range of cell processes, including access to DNA by transcription factors — the proteins that regulate gene expression. In other words, DNA must become unblocked to allow its information to be read and to produce messenger RNA (mRNA), which in turn must exit the nucleus and become translated into a protein product.
How that might occur — how DNA becomes more accessible to transcription factors — is currently an area of intense research scrutiny, including at UNC-Chapel Hill.
Strahl, Yi Zhang, assistant professor of biochemistry at the Lineberger Com-prehensive Cancer Center, and others have modified the older view of many scientists that histones play a passive role in chromosomal architecture, a view of histones as primarily structural, packaging DNA into chromatin fibers while having little to do with gene regulation.
Independently — Strahl working with yeast cells and Zhang with mammalian cells — the two are discovering that histones play a more dynamic role in chromatin, namely, its loosening or tightening.
The researchers’ attention is focused on histone methylation, the addition of a methyl group to lysine, one of the amino acids that comprise the tail region of histone molecules.
“We’ve known for three decades that histones can be methylated, but nobody knew the identity of any of the enzymes responsible for this methylation until two years ago,” Zhang says. That was when the first such enzyme was identified which specifically methylates histone H3 at lysine 9. Its presence there was linked to chromosome areas of gene silencing or inactivation.
Zhang’s lab has since identified the enzyme SET7, which specifically modifies lysine 4 on the histone H3 tail. This modification makes the chromatin structure more open so other proteins can access particular genes, Zhang says. Moreover, methylation of the same histone at lysine 4 and lysine 9 have opposite effects. Thus, according to Zhang, methylation at either site could determine either gene activation or gene silencing. Still, the situation is probably more complex than that. Among the possibilities, SET7 could have functioning partners yet unidentified, Zhang says. He recently reported discovering another two enzymes, and his lab is intensely studying their functions.
For his recent entry into histone modification, Strahl and former colleagues at the University of Virginia identified and characterized Set2, a novel histone that is responsible for methylating lysine 36 on the H3 tail. However, this modification helps to repress or silence gene transcription. Thus, Set2 might be “a coregulator of transcription” in the sense that it turns genes “off” instead of “on,” as in the case of SET7.
“During development, you have different sets of genes that are important for, say, limb formation, and when the limbs are completed, the genes responsible for them must be turned off,” Strahl says.
It may well be that methylation and other modifications are part of an emerging “histone code” of modifications that ultimately regulate gene expression. Strahl and his former mentor at the University of Virginia, David Allis, postulated such a code in a 2000 paper in the journal Nature. This code would be in addition to the now familiar genetic code of repeating As, Cs, Gs, and Ts of DNA nucleotide sequences. Through this histone code, differentially modified histone proteins could organize the genome into stretches of active and silent regions. Moreover, these regions would be inherited during cell division.
“We believe that methylation and other modifications that affect histone proteins, including acetylation and phosphorylation, are all dynamically involved and play critical roles in gene activation and deactivation at the appropriate times,” Strahl says.
This process, he explained, possibly could work by the ability of these modifications to bring in additional proteins that result in opening or closing of the chromatin molecule.
“Yi and I, as well as other labs, are at the frontier of understanding about the enzymes that are so important for the dynamic regulation of chromatin,” Strahl says. “Our findings add to our knowledge of a basic and very important process in human biology. They could offer new insight as to why certain genes in cancer are inappropriately expressed and how that might be corrected.”
Meanwhile, in Bill Marzluff’s sprawling and bustling Fordham Hall laboratory, the histone focus is even more basic than trying to tease out chromatin dynamics. “We work on the histone messenger RNA level — DNA’s blueprint for histone proteins — its regulation, processing, transport, and degradation,” graduate student Judy Erkmann says. Her work explores how histone mRNA is transported from the cell nucleus to the cytoplasm.
The focus on histone mRNA stems largely from the Marzluff team’s 1996 discovery and cloning of the stem-loop binding protein, SLBP. This unique protein is a major regulatory player in histone mRNA. It latches onto the looped tail of histone mRNA and signals the synthesis of histone proteins crucial to cell functioning during embryogenesis and throughout the organism’s life.
But SLBP does more than simply hitch a ride on a loop of nucleotides. It also doggedly performs a string of important duties after it takes the mRNA into a specific region of the nucleus. It interacts with other proteins to make sure the mRNA is properly processed into its final form. And then after helping get it out to the cytoplasm — the cell’s factory floor — SLBP remains bound to histone mRNA, making sure that its instructions are properly translated.
“A lot of people work on histones, but just a few groups in the world actually work on understanding how histone mRNA is synthesized and regulated,” says research assistant professor Zbigniew Dominski, whose own work focuses on how mRNA is processed and matures into a translatable message. “In this lab we cover all the different steps of histone mRNA message metabolism.”
“And that’s very important because the synthesis of histone proteins depends strictly on metabolism of the mRNA,” says Ricardo Sanchez, the lab’s newest Ph.D. recipient. Histone mRNA translation and regulation remain his major research interest.
And what if the metabolic regulatory process goes awry? Recent findings in Drosophila — fruit flies — by Robert Duronio, associate professor of biology, in collaboration with Marzluff, highlight the possibilities at the edge of life’s beginnings. Mutated or non-functioning SLBP is associated with failure to develop beyond the early embryo. A similar outcome may well apply to other multicellular creatures, including us.
Mapping Disease
From cystic fibrosis to space rats, proteomics is allowing researchers to uncover the secrets of disease.
Genome, transcriptome, and proteome are all fancy terms used by scientists to describe the separate populations of molecules that contribute to our beings, determining who we are. What color eyes we have, how fast we can run, if we are going to have diabetes — it’s all in our genes. With the completion of the human genome project came the hope that the molecular links to what makes us tick would emerge from the three billion As, Ts, Cs, and Gs that constitute our DNA. With this promise still unfulfilled, many scientists are turning to proteomics to study the products of our genes — proteins — and how they change structure, interact with each other, and give rise to disease.
According to Lee Graves, associate professor of pharmacology, proteomics is where the action is. “Proteins do everything,” Graves says. “They catalyze the metabolic functions of the cell, they break your glucose down to give you energy, they transmit signals from the outside of the cell to the inside of the cell.” While DNA is a static store of information, proteins are constantly changing, undergoing modifications, being depleted and degraded, acting differently in different tissues. Even though this complexity makes proteomics more challenging to study than genomics, it also confers the potential to greatly increase the understanding of human disease. Proteomics can be used to discover proteins that are associated with diseases and determine which of these can serve as novel targets for drug development or as biological markers of human disorders.
Proteomics enables scientists to study how proteins function in healthy cells as well as what goes wrong when disease strikes. Cystic fibrosis (CF), a common hereditary disease affecting approximately 30,000 children and adults in the United States, is just one of the many disorders that can potentially benefit from proteomics research. Even the identification of the cystic fibrosis gene over a dozen years ago has not led to an effective treatment. Although CF is caused by a defect in just one protein, known as cystic fibrosis transmembrane regulator (CFTR), it creates a variety of symptoms including faulty digestion due to a deficiency of pancreatic enzymes, difficulty in breathing due to mucus accumulation in airways, and excessive loss of salt in the sweat — all caused by one faulty protein.
“To try to understand why a protein works differently in a sweat duct than in an airway, we need to know all the partners, all the other proteins,” says Richard Boucher, director of the Cystic Fibrosis Center. According to Boucher, CF is caused not only by the absence of the CF protein’s function, but also by the absence of crucial interactions with other proteins in the cell. “There are ways of looking at those tissues and essentially identifying and cataloguing all the proteins in the tissues and then asking which ones are involved in CFTR,” he says. Boucher is using proteomics to identify the various parts and determine how they interact with the CF protein to form a functional cell. “Once we know how the system is wired together — that is, what wires or connections are missing because of the missing CFTR protein — then we can reconnect the system or jumpstart it with drugs,” Boucher says.
Muscle atrophy is another disorder that proteomics can help us understand. Diabetes, aging, and dystrophies such as muscular dystrophy all result in muscle wasting. Scientists at NASA are working with Graves to study the muscle atrophy experienced by astronauts after extended missions in space. “We are collaborating with a NASA group that has an animal model system to mimic weightlessness, and what we are trying to do is to apply modern methods of proteomics and mass spectrometry to analyze the changes,” Graves says. One of the well-observed responses to atrophy is a change in protein expression. By comparing protein levels in normal and atrophic muscle, scientists can profile these differences and determine which proteins are involved in muscle wasting. Tom Hilder, a graduate student in pharmacology, and Jun Han, a postdoctoral fellow, are both working in Graves’ lab to identify the important proteins so that therapies can be developed to hinder atrophy.
In addition to the discovery of novel drug targets, proteomics can also be used to identify biomarkers, which are specific profiles that indicate the severity of disease. “For cystic fibrosis, we really need to know how bad the infection and destruction is in the lung at any given time,” Boucher says. The current methods to assess the severity of disease — symptoms, chest X rays, and blood tests — are not very helpful, especially in children. Finding markers for lung inflammation and infection would help researchers decide when to initiate therapy and would help in clinical trials of new therapies. Margaret Leigh, professor of pediatrics, is collaborating with Boucher to locate these biomarkers by collecting one to two hundred serum samples from CF patients and then comparing the data with those from normal subjects. Using large 2-D gels and bioinformatics, Boucher can identify which of the 12,000 serum proteins vary between normal subjects and people with CF and of these, which ones increase or decrease with the severity of lung disease.
These biomarkers can be a great tool for pediatricians treating CF babies, who are born with normal lungs at birth and do not get their first infection until somewhere in the first few years of life. Pediatricians can use biomarkers to detect when that first infection occurs and initiate treatment with antibiotics to irradicate the infection before it begins to damage the lungs. The same biomarkers that track the severity of infection and inflammation of the lung can then be used to determine if a new drug ameliorates the patient’s condition. According to Boucher, this approach has already produced results with other disorders. “I think that there have been great successes with biomarkers and using proteomics approaches, predominately in cancer.”
Graves is working with other scientists to apply proteomics to cancer research. Carolina’s Lineberger Comprehensive Cancer Center has breast, prostate, and colorectal cancer projects in the works. “We can profile, or look at a patient’s sample, and say that the expression of this protein correlates with metastatic breast cancer or prostate cancer,” Graves says. “If we screen out these different patients, we can find something that is now expressed more highly in people that are showing advanced cancer versus those with early cancer or no cancer at all. As a pharmacologist or a biochemist, we can start to design drugs to attack the problem at the molecular level.”
Although proteomics entails fishing through thousands of proteins to find just a handful of useful candidates for drug targets or biomarkers, Graves doesn’t mind a bit. “There is a fair amount of fishing in science,” he says. “You just have to have the right net to make sure you catch something at the end of the day.”
Tiffany Heady was a student who formerly contributed to Endeavors.
Robin Arnette was a student who formerly contributed to Endeavors.