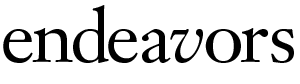Cell Talk
For years, when Ken Harden went home for the holidays, his Aunt Beulah would try to figure out what he did for a living. “What kind of research are you doing?” she would ask. “Cancer research?”
No, not exactly.
“Heart?”
Well, no.
“Then what, exactly?”
In a way, he was studying them all—every disease and disorder of vital interest to Aunt Beulah and her family and friends. Because Harden, professor of pharmacology, is one of an ensemble of Carolina scientists who have tuned their careers to the theme of cell signaling.
“All of a sudden,” Harden says, “we have become one of the best places in the country to do this kind of work.”
For Harden, Carolina jumped up the leaderboard when it landed several young stars who instantly meshed with the talent already in place. “If you were to tell anyone working in this field that we had snagged any one of these guys, they’d say, ‘Wow,’” Harden says. “And we have several wows.”
Why are the wows coming here? Why is a Monday-night session on cell signaling one of the hottest draws on campus, regularly packing in 40 to 60 faculty members, postdocs, and students who actually care passionately about G proteins and other exotica?
The short answer: this is the time and the place. Just as the revolution in cell-signaling research is about to explode, Carolina has been quietly assembling an arsenal of intellectual firepower. And now, in various combinations, the wows are setting off bombshells in journals such as Nature and Cell. These scientists believe, adamantly believe, that untangling the mysteries of cell signaling will be the next, necessary step toward treatments—or, dare we say it, cures—for cancer and many other diseases and disorders.
No, this is not science fiction. In clinical trials, patients with early phase chronic myelogenous leukemia are taking a pill that stops the disease in its tracks for at least 18 months. The pill contains a molecule that artfully intervenes in a cell-signaling process gone awry. And this remarkable result, scientists believe, is the first of many to come. With the sequencing of the human genome complete, and the computing power arriving to crunch the numbers, researchers and drug companies have scores of new molecules to investigate and many possible “targets” for treatments or cures in the pathways of cell signaling. So in the halls of biochemistry, pharmacology, medicine, radiation oncology, and other departments on campus, there is a sense of urgency, of dramas unfolding. Ideas pinball around from lab to lab. And everyone involved seems to run on adrenaline.
Harden, for one, is the kind of guy who will leap up from his chair and flip the lights on and off as he’s explaining a molecular switch. He will dash little red diagrams all over your notebook paper, trying to boil something on the order of War and Peace down to the level of “See Spot run.” Not all of the details sink in, but the fascination does. Here is a bit of the story, as we understand it, so far.
Loaded for Bear
In most cases, disease represents failure to communicate, not in the brain but down in the trenches, in cells. Sure, the brain thinks it’s in charge, and it does indeed command and control many processes necessary to life. But the signals in our bodies don’t stop at the ends of our nerves. They are everywhere, in every cell, moving messages along: go, stop, grow, change, die.
Cell signaling is the way a cell interprets information, not only from its environment but from its own genetic code. Cell signaling mediates our response to odors, to light, and to other kinds of stimulation. It also controls the enzymes of metabolism, controls how genetic information gets put to work (gene expression), and controls the cell’s shape and movement. Simply put, it controls the choices—the decisions cells make.
Many signals originate outside the cell and must negotiate their way through its defenses. Steroid hormones such as testosterone and estrogen melt their way through the cell membrane. But other kinds of signals have to dock onto the surface and push the right buttons. Consider a familiar example: adrenaline, that rush that comes with excitement or fear and gets our bodies ready for action.
“What happens when you encounter a grizzly bear or a shark?” Harden asks. “What do you want your body to do? You want your pupils to dilate. You want your bronchial smooth muscle to relax. You want your blood flow to be redistributed to tissues that are important for dealing with stressful situations. You want to break down glycogen so that you have glucose for energy.
“All of that occurs because there are cell-surface receptors that recognize adrenaline in a specific way, and these receptors coupled with two other proteins transduce the signal across the cell membrane to start cascading events inside the cell. So a lot of drugs that are used in everything from the treatment of glaucoma to all kinds of heart disease to the common cold interact with these sets of receptors for adrenaline, either by mimicking the effect of adrenaline or by blocking the effect of adrenaline.”
For years, Harden and his colleagues have been working out the signaling pathways associated with adrenaline and its cousins. Histamine, for instance, is involved in allergic reactions. Dopamine, a neurotransmitter, influences moods and addictions. And serotonin, which, among other things, constricts blood vessels, is associated with migraines and depression.
“All of these act in one way,” Harden says. “They all have a G-protein coupled receptor (see How to get through to a cell).”
Basically, the receptor is a biological switch. When it’s on, the signal flows. When it’s off, the signal stops. Many drugs or druglike compounds, natural or synthetic, work by turning these switches on or off. In fact, more than half of all drugs target the cell-signaling machinery. A well-publicized, recent example involves a molecule that signals the smooth-muscle relaxation that promotes blood flow. By tapping into its signaling pathway, pharmacologists and drug makers gave the world Viagra.
And then there’s caffeine, which gives us that wide-awake buzz. Normally, cell-signaling responses for adrenaline and serotonin and their like close down as enzymes called phosphodiesterases break down cyclic AMP, which is the “on” side of the switch. By inhibiting phosphodiesterases, caffeine keeps the switch open.
Each of the various enzymes involved in such pathways is a potential target, a place where a chemical nudge could, as Harden puts it, “restore a balance or reroute a hell-bound train.”
How Cancer Breaks the Suicide Pact
Not every signal originates outside the cell. Very often, the trouble that ends a life begins with a mistake, an internal error. Let’s say a random mutation strikes a molecule associated with cell growth. Suddenly, the cell is firing off the message to multiply, and unless another signal arrives to shut it down, the runaway cells become cancer.
This isn’t news. We’ve known this story for decades. But today, we know that the story was incomplete. For one thing, cells have the ability to correct some kinds of mistakes the way editors correct typos.
“We get damage to our DNA all the time,” says Shelton Earp, director of the Lineberger Comprehensive Cancer Center, “from our environment and from mistakes when DNA are replicating. There is a special set of genes that proofread the DNA. If they find mistakes, they can repair them. If they can’t repair them in a timely fashion, then they send a signal out to stop the cell cycle, paralyzing the cell at one stage so it will then have the time to repair that damage. And if that still doesn’t work, then it has to send a signal for the cell to die, to commit suicide.”
Cell suicide is not just a way of erasing mistakes. It can also be a necessary step in normal development. Earp holds his hand up, fingers together, mittenlike. “When you’re an embryo and your limbs are formed, you’re undifferentiated like this,” he says. “The way you get fingers is that the cells between the fingers die, and they die because of what is called ‘programmed cell death.’ In the genetic code there’s a program that leads to a set of signals that say, ‘Okay, these cells die,’ and you get digits.”
Since cell suicide is intimately involved with the arrangement of cells in our body plan, Earp wonders why it took researchers, himself included, so long to realize that cancer wasn’t just a problem of cell growth but was also a problem of cells’ failure to die.
“Some cancers do begin with a mutation in a cell-signaling molecule that signals for growth,” Earp says. “And when these mutations occur, they’re referred to as oncogenes. But that’s only a part of the picture. Most leukemias, for example, or lymphomas, are diseases that occur when a cell fails to differentiate or fails to die on time.”
Earp’s lab, working with Carolyn Sartor of radiation oncology and Ben Calvo of surgery, studies tyrosine kinases, a class of signaling molecules that, among other things, drives the growth of breast-cancer cells. By understanding the signaling pathways involved, the researchers find possible targets for drugs that could someday stop the cancer by flipping a biochemical switch. This kind of high-precision editing is very different from conventional cancer treatments. Chemotherapy is designed to attack, without discrimination, rapidly proliferating cells. Unfortunately, we also have normal cells that proliferate rapidy, such as the cells in our hair follicles and our bone-marrow, so chemotherapy attacks those, as well.
“If you look at cancer treatments before 1990,” Earp says, “you would say that almost none of the therapies, with the possible exception of hormonal therapies like Tamoxifen, involved cell signaling pathways. We’ve seen that change over the last ten years. But the next ten years will be the era in which cell signaling comes to the fore.”
The Information Explosion
One reason for this optimism is that the human-genome projects have cranked out an enormous index of genetic information, including many molecules that may be involved in signaling. Another is that new technologies are helping researchers mine meaning from this mountain of genetic information. For example, Chuck Perou, who recently came to Carolina from Stanford, helped pioneer a technique for rapidly measuring changes in the expression of genes, enabling researchers to “profile” cancers and their genetic origins.
But along with the opportunities, these advances have revealed enormous complications. Over the next decade, Carolina will invest at least $245 million in public and private funds to support studies in genomics. These will include work in bioinformatics, the intense computations required to make sense of the mountains of data from the human-genome projects and other sources.
“If you walk around the cancer center and look at the diagrams people are drawing on their boards, you’ll see that these signaling networks are in fact multiple networks, and they interact with each other,” Earp says. “So bioinformatics, or computational biology, is really trying to understand how these signaling mechanisms integrate in the cell.”
Channing Der, a professor of pharmacology who works with Earp at the cancer center, says that even if we understand a signaling mechanism in one cell type, it may not be the same in another. Der is a leading expert in the Ras oncoprotein, which regulates cell growth. Because human tumors frequently contain mutated Ras proteins, Der and others believe that the proteins are involved in cancer.
“The Ras oncoprotein we study is important in lung cancer, colon cancer, and pancreatic cancer,” Der says, “but how it contributes to the development of cancers in each of those organs may not be the same.”
Der’s lab, working closely with Adrienne Cox of radiation oncology, isolates specific molecules that show promise for cancer treatment, then works with drug companies to test the compounds on cultured cells and laboratory mice. Because these tests cannot reliably predict results in humans, the testing itself adds another level of complexity.
“When we decide a target is important and we make inhibitors against that target, they may work fantastically in cell-culture models or even in mouse models,” Der says. “But whether they will work in humans is a big question mark.”
Despite the complications, Der is convinced that the next generation of cancer treatments will arise from cell-signaling studies. “All you have to do is look at cancer survival rates for the last two decades and see that they haven’t improved dramatically,” Der says. “That’s because our conventional approaches, while useful, have simply exhausted themselves with regard to making a breakthrough, and so understanding signal transduction is the great hope. This just has to be how we’ll make inroads into solving the complexities of human disease.”
Picking the Right Pocket
Resolving the complexities of cell signaling is impossible for any one scientist or lab, so researchers collaborate, often with colleagues all over the world. But more and more frequently these days, Der is also finding partners here at home. Recently, he began working with John Sondek, a newcomer in pharmacology who specializes in studies of the structure of signaling molecules.
“Structure becomes critical when there are several molecules that are closely related,” Der says. “Let’s say you have three molecules, A, B, and C. A is the critical one, the one involved in cancer. So we want to make an inhibitor against A. But because B and C are similar, there’s a good possibility that the inhibitor might recognize B and C. So John Sondek’s lab might, for example, look at the structures of the inhibitor and its potential targets. A drug sort of fits a pocket. Each pocket is a little different. The more precisely you can design that drug so that it recognizes a specific pocket, the more likely you’re going to be able to make a drug to target protein A but not B and C.”
Sondek and David Siderovski, another newcomer in pharmacology, are in demand as research partners because their work bridges various kinds of cell-signaling studies, engaging the team in what Ken Harden calls “crosstalk.”
Until recently, the signaling pathways explored by cancer researchers like Der and Earp—pathways having to do with cell growth and regulation—didn’t seem directly involved with the hormone-and-neurotransmitter signaling that Harden and others pursued. Sure, the two groups have always spoken the same language, but now they are part of the same conversation.
“We’re finding that these two areas connect in a lot of very interesting ways,” Harden says. “And these new guys, John Sondek and David Siderovski, are right in the middle of the crosstalk. So is Henrik Dohlman, recently recruited from Yale by David Lee, the chair of biochemistry.”
The Case of the Anxious Knockout
Dohlman, Siderovski, and others independently discovered a class of signaling molecules known as RGS proteins, which help regulate the on-off switches in various signaling pathways. These RGS proteins are part of the crosstalk. They are also evidence of just how deep the mysteries of cell signaling can run.
Take the case of RGS2, for instance. Siderovski has found that RGS2 affects the immune system by telling T cells how fast to launch a defense against invaders. When he studied mice in which RGS2 had been genetically “knocked out,” Siderovski found that their immune responses slowed dramatically.
This in itself was intriguing. But then the results took an astonishing turn. The knockout mice, which looked normal and displayed normal cognitive and motor skills, were remarkably anxious and fearful. They avoided light, backed down from confrontation, and were bullied by their normal brothers. Why would a protein involved in the regulation of immune response also affect fear and aggressive behavior?
“We don’t know,” Siderovski says. “We have seen that RGS2 controls an anxiety-aggression circuit in the brain, but now we need to swim upstream and find out what receptor is firing too often in the brains of these knockout mice. If we can piece together the molecular reasons for this behavioral deficit, we might have another avenue to develop new antianxiety drugs.”
Reading the Tea Leaves
Most likely, this quest will begin where his others have—with Siderovski staring at protein sequences on a computer screen, trying, as he puts it, to read the tea leaves. “I always say that my hypotheses are generated in silicon, rather than in the test tube,” he says. “And then we’ll go to the bench, purify the proteins, put them in neurons, make a knockout mouse, and do what we have to do to test our hypotheses. But they all start with this new explosion of bioinformatics.”
So Siderovski has been cast as the data guy, Sondek as the structure guy. And both of them believe that Rudy Juliano, their chair in pharmacology, had a design, a strategy in mind when he recruited them. More than individual talents, they work as complements, a team greater than the sum of its parts.
Siderovski has a favorite example. He and Sondek had been using a computing technique to delve into the genome, trying to find binding partners—places where signaling molecules connect—for two proteins they had been studying. “And there were hits,” Siderovski recalls. “But they were reported as being statistically insignificant, as being in the noise. So John and I went back to the literature, and we jogged back and forth. And this is the beauty of this collaborative place: we kept talking about it. And we finally teased out a deduction that both of our molecules bind to the Ras molecule, which is, as Channing Der will tell you, a central oncogene.”
It was a hunch that paid off, and it was one they might not have been free to follow, somewhere else. Siderovski, who worked several years in industry, jumped back into academia because, he says, he felt passionate about RGS proteins and wanted to pursue them on his own terms.
“In industry, the shareholders can’t wait,” Siderovski says. “But here, we can have a longer-term vision. If we work on basic research for five years, we might come up with ten drug targets, each of which could save lives and generate a billion dollars. Because here, the National Institutes of Health and the people of North Carolina are making the investment.”
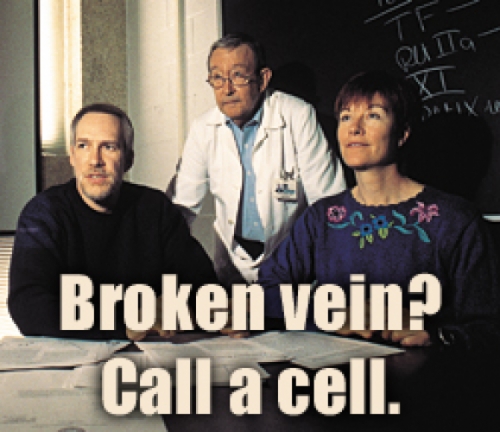
Broken vein? Call a cell.
You’re walking down the street, and a few blood vessels break in, say, your knees. Within minutes, tiny blood clots form—but only in the precise area of those breaks. This process happens many times a day.
If your body doesn’t continuously make these tiny clots, you have hemophilia and bleed uncontrollably. Even more common is thrombosis—when clots spread too far or occur in the wrong places—which often leads to heart attacks.
How does your blood know when to flow and when to clot? The answer, say some researchers, is cells.
“Cells talk to one another,” says Harold Roberts, professor of medicine. “It’s like they’ve got a telephone line between them.” The cells do their “talking” through receptors, which are proteins that will bind only to another specific protein. When that binding happens, a change occurs in the cell. That change is known as a signal.
Roberts and his collaborators, Mac Monroe and Maureane Hoffman, have developed a textbook-changing model of how cells control blood clotting. Other Carolina researchers are studying the receptors involved in that control.
Cells haven’t always gotten this much attention. For 50 years, blood clotting has been thought to be controlled by 12 proteins called clotting factors. (For simplicity’s sake, most of the factors are referred to by a roman numeral.) The factors have been studied intensely, and researchers know for example, that if you’re missing factor VIII or IX, you have hemophilia.
Cells, on the other hand, have been widely thought of as just “bags of lipid,” says Monroe, assistant professor of medicine. Lipids, which make up the membranes of cells, are molecules made of a water-soluble part and chains of fat. One lipid in particular, phosphatidylserine (PS), regulates one of the last steps in clotting. Some coagulation researchers still believe that cells are good for only one thing: to provide a surface containing PS where the clotting factors can assemble. And many researchers don’t use whole cells in their experiments. Instead they use synthetic lipid vesicles (small liquid-filled sacs) that contain PS.
“Cells are really messy,” Monroe says. “You have to grow them or isolate them from someone.” The thinking has often been if the synthetic vesicles serve the same purpose, then why bother with cells?
Monroe can understand that reasoning because he’s a biochemist, which means he is, in his own words, a control freak—he likes a nice, clean experiment that gives reliable results. But he also knows that in our bodies things aren’t always that simple. And if Monroe forgets, Roberts, a physician who regularly sees hemophilic and thrombotic patients, will remind him. And then there’s Maureane Hoffman. Now director of the Coagulation Laboratories and blood bank at the Durham Veterans Affairs Medical Center, Hoffman used to do research at Carolina and first got to know Roberts and Monroe when she began giving them platelets that were a waste product of her work. Of the three researchers, she knows the most about how cells work, and she makes sure the group’s experiments take cells’ properties into account.
The three believe that the traditional model of blood clotting tells us a lot about how the clotting factors react with each other but that it downplays the role of cells too much. For one thing, the traditional model of coagulation doesn’t always jibe with what happens in our bodies. It doesn’t explain how clotting occurs in only the tiny place where a break occurs. Nor does it explain why the lack of factor IX causes hemophilia in humans. In a test tube with synthetic lipid vesicles, blood will clot without factor IX.
Roberts, Hoffman, and Monroe work to design experiments that combine the control of biochemistry with the complexity of the human body. In their test tubes, they use whole cells such as human platelets, as well as the clotting factors. But not just one or two in isolation—a whole bunch together. “We know some of the important players, but biochemistry hasn’t looked at them in a complex,” Hoffman says.
After nine years of research, the team has developed a model of blood coagulation that has begun appearing in new editions of Williams Hematology and the Oxford Textbook of Medicine. Instead of depicting coagulation as a “cascade” of reactions among the clotting factors, the new model shows clotting happening in three stages, all controlled by cells.
The first stage, initiation, happens on a cell that has tissue factor on its surface. Tissue factor is a protein that stays on the membrane of a cell at all times. Your blood normally never touches tissue factor—unless a blood vessel breaks. Then plasma in the blood is exposed to tissue-factor-bearing cells, and the tissue factor binds to one of the clotting factors in the plasma, factor VII, which becomes activated or turned on. (The activated form of a factor is noted with a lowercase “a,” as in VIIa.)
Tissue factor and factor VIIa are what are known as an enzyme and a cofactor. An enzyme speeds up a chemical reaction, and its cofactor helps the enzyme do its job faster. So, bound together, the tissue factor and factor VIIa activate factor IX and X (creating IXa and Xa). These combine with other factors, continuing until factor Xa combines with factor Va, which changes an enzyme called prothrombin into its active form, thrombin. Thrombin is one of the compounds that starts the clotting process. But at this stage just a small amount is produced—not enough to cause a clot.
It is enough thrombin, though, to begin activating platelets that have been circulating in the blood in an inactive state. This is the second stage, called amplification, in which the action moves to the platelet surface. Once activated, platelets release their internal stores—sort of turn themselves inside out. One of the things that’s released is a small molecule called adenosine diphosphate (ADP), which will bind to a receptor on another platelet and activate it. “So the activated platelet is sending an inactivated platelet a signal—‘stick down here,’” Monroe says. The activated platelets go directly to the site of the bleeding, where they form a sticky mass—the first soft clot that you see after you cut yourself.
In the final stage, propagation, activated factors combine with their cofactors and bind to sites on the surfaces of platelets. One crucial part of this stage is when IXa and VIIIa combine to form Xa, which helps produce a large burst of thrombin. The large amount of thrombin sends signals that turn a factor called fibrinogen into its active form, fibrin, which weaves through the platelet clump to form a scab.
The model provides clues as to why people can’t clot without factor IX but blood in a test tube can. In the last stage, activated factor IX helps make the crucial Xa on the platelet surface. But factor Xa is also produced earlier, by factor VIIa and tissue factor. So why can’t this Xa compensate for the Xa normally made by IXa? In a test tube, it can. But in a person, the first Xa is produced on a tissue-factor-bearing cell. For the Xa to promote clotting, it must move from the tissue-factor-bearing cell to the platelet surface. But inhibitors in the plasma probably stop it before it can get there, Roberts says.
At a more minute level, Leslie Parise, professor of pharmacology, studies how thrombin signals the platelet to bind to fibrinogen. When thrombin or ADP activate their receptors, some of the signals created inside the platelet turn on a glycoprotein called GP IIb-IIIa. It is GP IIb-IIIa that actually binds to fibrinogen, forming that first soft clot at a site of blood vessel injury, Parise says. Her lab has identified a new protein that they think may be involved in regulating the activation of the receptor for GP IIb-IIIa. “The platelet tightly regulates the activation of this glycoprotein,” Parise says, “because if it was activated, our blood would clot and we’d be having heart attacks all the time.”
Roberts’ group thinks that clotting is also kept in check by endothelial cells (those that line the inside of the blood vessels). Normally, platelets don’t stick to endothelial cells because these cells produce an enzyme that degrades ADP. So if a platelet accidentally gets activated, this enzyme “chews up” the ADP so that it can’t signal more platelets to activate, which could cause a clot that could block blood flow to the heart or lungs.
As many details as are known about this process, there as just as many that are a mystery. For instance, scientists know that the binding sites for clotting factors exist on the platelet surface because they can measure the sites’ activity, Monroe says. But no one has identified the protein structures of most of the sites, much less sequenced them or cloned them. And researchers don’t know yet whether these sites are true receptors that signal changes inside the cell or simply binding sites (click here for a diagram).
As these researchers learn more about how cells control clotting, they hope they will learn more about how we can control it. “Right now, your control mechanisms are keeping you alive,” Roberts says. “If we can understand the normal mechanism, then we can figure out what goes wrong, what is it that causes disease.”
Platelets—cells that form the first sticky plug of a blood clot—are activated by an enzyme called thrombin. When one platelet gets activated, it sends a signal to another platelet to activate, and so on. If platelets activate when they’re not supposed to, you can develop a dangerous clotting of the veins. To learn how the body keeps clotting under control, JoAnn Trejo, assistant professor of pharmacology, studies the receptor for thrombin on a platelet’s surface.
Once thrombin’s receptors have done their job, they are removed from the cell surface to a compartment within the cell called the lysosome, which degrades the receptor (“chops” it into its single amino acids). This degrading is an unusual aspect of thrombin’s receptors. Many other types of receptors are recycled—they move to the inside of the cell, are modified slightly, and then return to the cell surface in an “off” state, ready to signal again.
“To get new receptors for thrombin back on the cell surface, the cell has to synthesize more receptors, and that costs a lot of energy,” Trejo says. “So it’s not a very efficient system.” But it’s an important one. Receptors for thrombin have to be degraded, Trejo says, because if they return to the surface, they can continue to signal.
But Trejo has found that thrombin’s receptors take the same pathway to get inside the cell as receptors that are recycled. So she thinks that there must be something particular about thrombin’s receptor that says to the cell “degrade me.” Her next step is figuring out how the receptor does that.
A Crazy Idea
Barry Lentz, professor of biochemistry and biophysics, doesn’t sleep much at night. He’s got about eight papers to write and publish, and he wants to do it fast. Lentz has spent years studying just one element of blood clotting—phosphatidylserine (PS). And now he’s found something unusual, something that he believes will shake scientists up. But it won’t do that unless they read about it.
PS is a lipid—a long molecule with a water-loving “head” and a greasy, water-hating “tail.” In water, lipids organize themselves into sheets so that the water-soluble heads face the outside of the sheet and the greasy, water-insoluble tails face the inside. This “lipid bilayer” is what forms cell membranes.
Most researchers have believed that the membranes of platelets help regulate the production of thrombin, which is an enzyme that helps your blood to clot at a wound. While these researchers have recognized that PS plays a role in this process, they have focused on the importance of the cell membrane surface. But work in Lentz’s lab shows that the water-soluble part of PS, without the cell membrane, enhances thrombin production.
Lentz had suspected that PS has a binding site all its own on Factor Xa, one of the proteins that helps make thrombin. But this was hard to prove with PS in a membrane—with the whole membrane involved, who could tell what exactly was binding to the Xa? Then Lentz asked a postdoc, Vishwanath Koppaka, now an assistant professor at the University of Pennsylvania, to work with a special soluble form of PS that has a very short, greasy tail. Koppaka found that the soluble PS would bind to Factor Xa, same as normal PS. And using fluorescence, the researchers were able to characterize the binding site. “People have thought that it was just the membrane that was important, that there wasn’t a specific interaction between the phosphatidylserine and Xa,” Lentz says. “But we’ve shown that there is.”
Lentz’s lab has since shown that another molecule, factor Va, also has binding sites for PS. Lentz now thinks that PS is a message molecule that signals clot formation at a wound, which may make it a target molecule for drug design.
Even once Lentz gets his papers published, this may not be a popular idea. “I am sure Barry is right,” says Mac Monroe, assistant professor of medicine. “But for a while it may be hard for him.” Since it’s been a “given” that all lipid actions occur as part of a membrane structure, the tendency may be to think that “anyone who thinks the water-soluble part of lipid has activity independent of its membrane association is ‘crazy,’” Monroe says. “But Barry is ‘crazy,’ and he is right.”